Scientists Uncover How Bacteria Repurpose Ancient Immune Systems to Fuel Antimicrobial Resistance Spread
Scientists have made a significant breakthrough in understanding the intricate mechanisms by which bacteria exchange genetic material, particularly genes that confer antimicrobial resistance (AMR). This critical discovery, stemming from extensive research at the John Innes Centre, sheds new light on the function of unusual, virus-like particles known as gene transfer agents (GTAs) and their pivotal role in facilitating this exchange, a process that underpins a growing global health crisis.
The Silent Pandemic: Understanding Antimicrobial Resistance
Antimicrobial resistance (AMR) is widely recognized as one of the most pressing global health threats of the 21st century. It occurs when microorganisms—bacteria, viruses, fungi, and parasites—evolve to withstand the effects of antimicrobial drugs, making infections harder to treat and increasing the risk of disease spread, severe illness, and death. The World Health Organization (WHO) estimates that bacterial AMR alone contributed to nearly 5 million deaths in 2019, with 1.27 million directly attributable to drug-resistant infections. Without urgent action, projections indicate that AMR could cause 10 million deaths annually by 2050, surpassing cancer as a leading cause of mortality and incurring a cumulative economic cost of over $100 trillion.
The rapid rise of AMR is multifactorial, driven by the overuse and misuse of antibiotics in humans, livestock, and agriculture, coupled with a dearth of new antimicrobial drug development. Bacteria, however, are not passive recipients of these pressures; they are incredibly adaptable organisms. A key aspect of their adaptability, and a major driver of AMR, is their ability to share genetic information through a process called horizontal gene transfer (HGT). Unlike vertical gene transfer, where genetic material is passed from parent to offspring, HGT allows bacteria to acquire new traits, including antibiotic resistance genes, from unrelated individuals or even different species within a single generation. This rapid exchange transforms bacteria into formidable adversaries, capable of quickly evolving defenses against our most potent medicines.
Gene Transfer Agents: Domesticated Viruses as Genetic Couriers
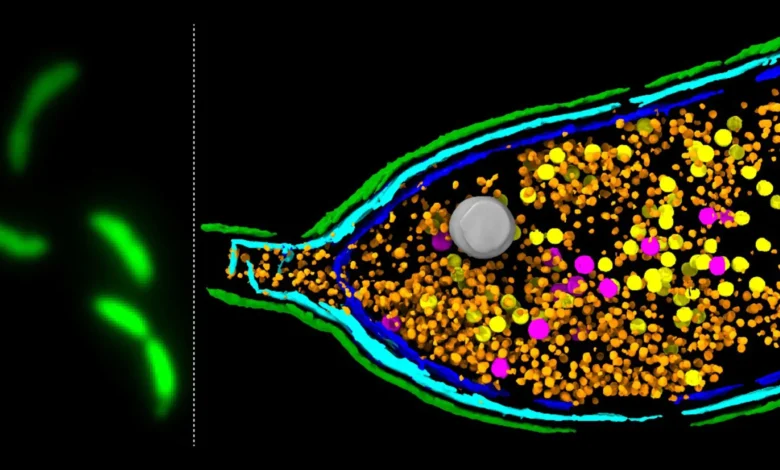
The John Innes Centre team’s research focuses on a particularly intriguing form of HGT mediated by gene transfer agents (GTAs). These particles bear a striking resemblance to bacteriophages—viruses that infect bacteria—but they are not the harmful invaders they once were. Instead, GTAs represent a fascinating evolutionary adaptation: they are remnants of ancient viruses that bacteria have, over millennia, domesticated and brought under their own control. Rather than replicating themselves or destroying their host to spread viral genes, GTAs function as tiny, specialized delivery vehicles for bacterial DNA.
The concept of GTAs first emerged from observations in the 1970s with Rhodobacter capsulatus, where researchers noted the transfer of bacterial genes by phage-like particles. Since then, GTAs have been identified in a diverse range of bacterial and archaeal species, suggesting a widespread and ancient evolutionary phenomenon. These particles package random fragments of their host cell’s DNA and then release them, allowing other nearby bacteria to take up these genetic "parcels." This "selfless" act of genetic sharing is a powerful mechanism for disseminating beneficial traits throughout bacterial populations, including, critically, genes that confer resistance to antibiotics. While other forms of HGT exist—such as conjugation (direct cell-to-cell transfer via plasmids) and transformation (uptake of free DNA from the environment)—GTAs represent a unique, self-produced mechanism that actively packages and delivers genetic material.
Unraveling the Mystery of GTA Release: The Host Cell Lysis Puzzle
A crucial step in the life cycle of GTAs, and therefore in the process of horizontal gene transfer, is host cell lysis. This is the controlled breaking open of the bacterial cell that produced the GTAs, allowing the DNA-packed particles to be released into the environment where they can then be taken up by other bacteria. Despite the recognized importance of GTAs, the precise molecular mechanisms by which these particles escaped from their host cells remained largely unknown until this recent study. Understanding this process is paramount, as it represents a potential choke point in the spread of resistance.
The researchers, led by a collaborative effort between the John Innes Centre, the University of York, and the Rowland Institute at Harvard, embarked on a detailed investigation using the model bacterium Caulobacter crescentus. Caulobacter crescentus is a well-established model organism in microbiology, prized for its relatively simple life cycle, genetic tractability, and its unique dimorphic cell division, making it ideal for studying fundamental bacterial processes. By employing a sophisticated deep sequencing-based screening method, the team systematically analyzed the genes involved in GTA activity within Caulobacter crescentus, aiming to pinpoint the genetic components responsible for host cell lysis.
The LypABC System: A Central Control Hub for Gene Transfer
The deep sequencing approach, a powerful technique that allows for the rapid and comprehensive analysis of genetic material, proved instrumental in identifying a critical three-gene system. This system, dubbed LypABC, encodes a set of bacterial proteins that the researchers found to be directly responsible for controlling GTA-mediated cell lysis.
To confirm the function of LypABC, the team performed elegant genetic experiments. When the lypABC genes were precisely deleted from the Caulobacter crescentus genome, the bacterial cells lost their ability to break open and release GTA particles. This demonstrated that LypABC was essential for the lysis process. Conversely, when the researchers engineered bacteria to "overactivate" or overexpress the lypABC genes, they observed a significantly higher proportion of cells undergoing lysis. These compelling results firmly established LypABC as a central control hub, dictating the release of GTAs from their bacterial hosts. The precision of this control is a testament to bacterial evolutionary fine-tuning, ensuring that genetic material is disseminated effectively.
A Repurposed Immune System: Evolutionary Ingenuity at Play
One of the most surprising and significant findings of the study was the evolutionary origin of the LypABC system. Molecular analysis revealed that LypABC closely resembles a bacterial anti-phage immune system. Bacterial immune systems, much like those in higher organisms, are designed to detect and neutralize threats, particularly viral infections from bacteriophages. These systems often contain specific protein components, such as CARD-NLR-like domains (as mentioned in the original paper’s title, "A bacterial CARD-NLR-like immune system controls the release of gene transfer agents"), which are typically associated with defense mechanisms against invading viruses.
The discovery that LypABC, a system facilitating bacterial gene transfer, shares such striking homology with an immune system is a remarkable example of evolutionary repurposing. It suggests that, over evolutionary time, bacteria have taken an existing biological system originally developed for defense against external threats and adapted it to serve an entirely new, self-beneficial function: promoting the efficient release of GTAs to share genetic material. This highlights the incredible plasticity and efficiency of bacterial genomes, where existing machinery can be co-opted and rewired for novel biological roles. This "exaptation," where a trait evolved for one purpose is then used for another, is a common theme in evolution but rarely observed with such clarity in the context of fundamental bacterial processes.
Tight Regulation: A Balancing Act for Bacterial Survival
The researchers also uncovered another layer of complexity: the LypABC system is under strict regulatory control. They identified a specific regulatory protein essential for maintaining this tight control over both the activation of GTAs and the subsequent lysis of the host cell. This stringent regulation is not merely an evolutionary quirk; it is critical for the survival of the bacterial population. The process of host cell lysis, while necessary for GTA release, is inherently destructive to the individual bacterium. Improper or unregulated activation of LypABC can be highly toxic, leading to widespread and potentially detrimental cell death within the bacterial community. Therefore, bacteria must carefully balance the benefits of gene transfer with the costs of sacrificing individual cells, ensuring that lysis occurs only when it is advantageous and controlled. This delicate equilibrium underscores the sophisticated genetic programming within bacterial populations.
Expert Perspectives and Future Directions in the Fight Against AMR
Dr. Emma Banks, the first author of the study and a Royal Commission for the Exhibition of 1851 Research Fellow, articulated the significance of these findings: "What’s particularly interesting is that LypABC looks like an immune system, yet bacteria are using it to release GTA particles. It suggests that immune systems can be repurposed to help bacteria share DNA with each other – a process that can contribute to the spread of antibiotic resistance." Her statement encapsulates the novelty of the discovery and its immediate relevance to the AMR crisis. The insights gained from this research are not merely academic; they provide a deeper, more granular understanding of how resistance genes disseminate, which is a crucial prerequisite for developing effective countermeasures.
The implications of this work are far-reaching. By revealing the precise molecular machinery governing GTA release, the study opens up new avenues for potential therapeutic interventions. If scientists can find ways to inhibit the LypABC system, or disrupt its regulatory control, they might be able to block the spread of resistance genes, thereby preserving the efficacy of existing antibiotics. This could involve developing novel compounds that specifically target LypABC proteins, or strategies that interfere with their activation. Furthermore, understanding the evolutionary repurposing of immune systems could offer broader insights into bacterial adaptability and evolution, potentially uncovering other hidden mechanisms that contribute to their survival and pathogenicity.
The immediate next steps for the research team involve delving deeper into the mechanics of the LypABC system. Specifically, they aim to uncover precisely how the LypABC control hub is activated and the detailed molecular cascade that leads to the rupture of bacterial cells and the release of GTA particles. This will involve investigating the signals that trigger LypABC activity, the structural changes it induces in the bacterial cell wall, and the interplay with other cellular components. Such detailed mechanistic understanding is essential for translating these fundamental discoveries into practical strategies for combating AMR.
In conclusion, this groundbreaking research from the John Innes Centre and its collaborators has illuminated a fundamental biological process at the heart of bacterial evolution and adaptation. By revealing how bacteria repurpose ancient immune systems to facilitate the spread of genes, including those that confer antibiotic resistance, the study offers vital new clues in the urgent global fight against antimicrobial resistance. It underscores the ingenuity of bacteria as evolutionary survivors and provides a critical foundation for developing innovative strategies to protect public health from the looming threat of drug-resistant infections.